Looking for Help with Assignments?
-RUCats has 80 hours of tutoring available online and in-person. Check the tutoring tab in Canvas!
-Instructors and Lead TAs have a combined 10 hours of Office Hours, open to all sections. See times/locations here.
-Piazza (found in the Canvas sidebar) provides fast support from Course Staff and other Students. Be sure to search to see if someone has asked a similar question!
-If you need a computer to complete work on, iLab machines can be found in the CAVE (Hill 252) and surrounding rooms.
DNA Profiles – 110 course points
The purpose of this assignment is to practice using Binary Search Trees to analyze DNA.
This assignment will take LONGER to complete than any of the labs.
Start your assignment early! You will need time to understand this assignment and any questions it may present.
Refer to our Programming Assignments FAQ for instructions on how to install VSCode, how to use the command line and how to submit your assignments.
The assignment has two components:
Coding (107 points) submitted through Autolab.
Reflection (3 points) submitted through a form.
Submit the reflection AFTER you have completed the coding component.
Be sure to sign in with your RU credentials! (netid@scarletmail.rutgers.edu)
You cannot resubmit reflections, but you can edit your responses before the deadline by clicking the Google Form link, signing in with your NetID, and selecting “Edit your response.”
Overview
You’ve been tasked with identifying who a DNA sequence may belong to based on an unknown person’s DNA sequence and a DNA database you have. A person’s DNA has pairs of chromosomes; we will focus on a single pair of the same chromosome to identify people whose samples we want to investigate further.
99.9% of our DNA is the same as other humans. Our goal is to focus on the 0.1% of DNA that varies between humans – these are sections of “noncoding DNA” that are in between genes and don’t provide instructions for making proteins. Short Tandem Repeats, or STRs, are in noncoding DNA and are short sets of bases that are repeated across a sequence.
In this assignment, you will work with simple DNA sequences with 2-5 STRs. We will identify the number of times a set of STRs is repeated within a sequence to find out who a pair of unknown DNA sequences may belong to in a DNA database.
Here’s a diagram that shows how we are using nodes to store Profiles:

Implementation
Overview of files provided
The Profile class allows you to create instances of Profiles; it stores information such as the full name of the profile (used as the key in the BST), whether the profile is marked as a profile of interest as well as an array of STRs (ie. which STRs appear in a person’s DNA and how many times). Do not edit or submit to Autolab.
The DNAProfiles class contains methods that manipulate BSTs in order to conduct STR analysis. Edit the empty methods with your solution, but DO NOT edit the provided ones or the method signatures of any method. This is the file you submit.
The Driver class is used to test your methods interactively. Follow the prompts in the terminal to use the driver. It is provided only to help you test your code. It is not submitted and it is not used to grade your code. Do not submit to Autolab.
The Queue class is provided from the textbook; it contains enqueue and dequeue operations. Do not edit or submit to Autolab.
StdIn and StdOut are libraries that handle input and output. Do not edit these classes.
Multiple input files are included; they store data handling DNA profiles and the unknown chromosome parts. You are welcome to create your own input files, as they will not be submitted to Autolab.
DNAProfiles.java
- DO NOT add new import statements.
- DO NOT change any of the method’s signatures.
Working with BSTNode<> objects
For assignments/labs in CS112, we will use a cs112.jar file containing useful classes. This includes the BSTNode<> class, which implements a binary tree node. You’ve previously worked with the LLNode<> class to create linked lists. Many aspects of the BSTNode are similar:
The BSTNode class takes in a generic type, which means it can store any type of data. In this lab you will be working with nodes that have Profile objects as their data – to have nodes with City data we can use BSTNode<Profile>
Making a node: BSTNode<T> newNode = new BSTNode<T>(data) — replace T with a type
Note that the data must be passed in for the constructor, and the data a node holds can not be changed later (aka it is immutable). The data may not be null.
The Profile object within a BSTNode is mutable, but the BSTNode itself is not mutable. You can access the Profile object using getData() and modify it from there if needed.
Methods:
getLeft() – returns the left child of the node
setLeft(BSTNode<T> left) – sets the left child of the node
getRight() – returns the right child of the node
setRight(BSTNode<T> right) – sets the right child of the node
getData() returns the data object a node holds
The DNAProfile class has the following instance variables:
BSTNode<Profile> rootNode – the root of our binary search tree
unknownSequence1 and unknownSequence2 – the 2 unknown sequences we will investigate, these will be read from the input file for you in a provided method
Methods to be implemented by you:
assembleTree (provided)
This method reads a DNA database from an input file as well as two chromosomes belonging to an unknown person. It then calls the following methods to insert profiles into a BST rooted at rootNode.
This method reads the full name of a person (“Last, First” including the space in between, case-sensitive).
This method calls createSingleProfile to read and return a single profile from the input file.
Then, it calls insertPerson to create a BSTNode containing the person’s name and newly built profile and insert the profile into rootNode.
Recall that:
The reference variable rootNode is the root of the BST.
Each BSTNode has a profile associated with it.
The BST then holds all the Profiles that are in the database.
Do not update this method, it has been implemented for you. IMPLEMENT the first two methods (createSingleProfile and insertPerson).
1. createSingleProfile
Given a single person’s data in an input file, read all information and return a new Profile containing the person’s information.
You have been provided with input files to test this method. Each input file is formatted as follows.
One line containing a string with the person of interest’s FIRST sequence [DONE for you in assembleTree]
One line containing a string the person of interest’s SECOND sequence [DONE for you in assembleTree]
One line containing the number of people in the database, say p [DONE for you in assembleTree]
For each of p people:
One line contains first name, last name, and number of STRs (say s), space separated. Names are read for you in assembleTree!
s lines containing an STR name and number of occurrences in that person’s DNA, space separated
To complete this method, do the following:
For a single person in the database:
Read the number of STRs (say s) associated with the person. You will create an array of STR objects of length s.
For each STR:
Read the STR name and number of occurrences.
Create a new STR object, and add it to the next open space in the array.
Create a Profile object using the data you just read, and return this profile.
Here is a diagram that illustrates a Profile object.
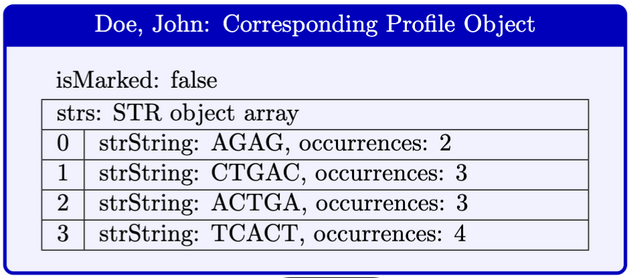
Use the StdIn library to read from a file:
StdIn.setFile(filename) opens a file to be read.
StdIn.readInt() reads the next integer value from the opened file (whether the value is in the current line or in the next line).
StdIn.readString() reads the next String value from the opened file.
Here is the expected output when testing createSingleProfile using the driver on input1.in. The driver calls the method on all people but does not update the unknown sequence instance variables. It also skips over names.
We expect STRs to be in the SAME ORDER as the input files.

2. insertPerson
Given a Profile object passed through parameters, add a node containing this name profile to the BST rooted at rootNode. We will be using the full name as the keys for insertion, following lexicographic order. Remember that rootNode is a reference to the root node of the BST.
Follow the BST insertion algorithm as discussed in class to insert Profiles into the BST.
Names are unique, nobody will share the same name.
Follow BST ordering when inserting a profile. Use the compareTo method on the keys (full names) for comparisons. BSTNodes with names greater than a node go to the right of the node, otherwise they will go to the left.
Here is a diagram that illustrates the correct output for the entire structure in input1.in.
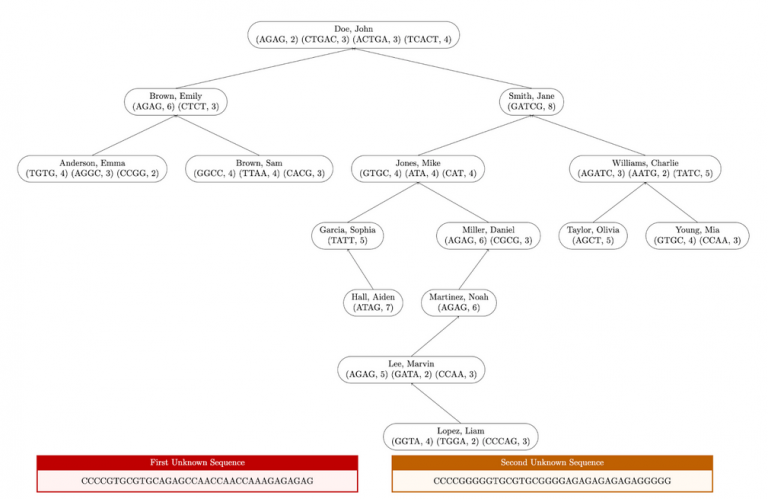
Here is the expected output when testing this method on input1.in. Test this method by calling assembleTree in the driver, it requires createSingleProfile to be completed.

Early submission. Implement the first two methods for extra credit: createSingleProfile() and insertPerson().
numberOfOccurrences (provided)
This method takes in a sequence and STR. It returns the number of times an STR appears in the parameter sequence. This method is provided, do not edit it.
-
You will call this method later in markProfilesOfInterest.
3. searchByName
Given a formatted full name as a string, recursively search the BST rooted at rootNode to try and find the profile it is associated with. If we find the profile associated with this name, we return the profile object, otherwise if it is not found we return null.
This method must be recursive, you must fill in the recursive helper method called searchByName with 2 parameters: BSTNode<Profile> currentNode, String fullName. The first searchByName is provided to you and calls the helper method which you must implement.
Use the compareTo() method on the full names of the BSTNodes to help determine which subtree your recursive call should lead you towards next (left, right, or found if compareTo() returns 0).
To test this method using the driver, first run assembleTree(), then run searchByName() and input a full name in the format “Last, First” (case sensitive).
Example with input1.in:
Enter FULL name of person to search for (Last, First – case-sensitive) => Lee, Marvin
Lee, Marvin found!
(Lee, Marvin) Marked: false
STRs:
AGAG, occurrences = 5
GATA, occurrences = 2
CCAA, occurrences = 3
If the full name we entered is not present in the BST, the Driver will tell you so as well:
Enter FULL name of person to search for (Last, First – case-sensitive) => Robinson, Curtis
Robinson, Curtis not found. Search returned null.
4. markProfilesOfInterest
This method marks profiles where for all STRs in a profile, most (at least half) of the STR occurrences in a profile match the occurrences of the same STR in the first and second unknown sequences combined. This is a simplified matching algorithm, and in reality the identification process is more complex.
-
Traverse the BST rooted at rootNode to investigate whether each profile should be marked.
-
For a single profile, we must mark it if at least half of its STRs occur the same amount of times in both the profile, and the first+second unknown sequence (the instance variables). Remember each profile has an “isMarked” boolean which is originally false when the profile is created.
-
We determine half of STRs by rounding UP to the nearest whole number if there is a remainder. For instance, if there are 3 STRs in a profile, half of 3 is 1.5, which rounds up to 2.
Hints:
-
Use the provided numberOfOccurrences method to check the number of times an STR is repeated in a single sequence.
-
Look at the STR.java class to see the STR object and the provided getter methods it contains.
In the following diagram, nodes with marked profiles are red and bold. Unmarked profiles are uncolored and unbolded. Profile STRs that appear in either unknown sequence are highlighted.
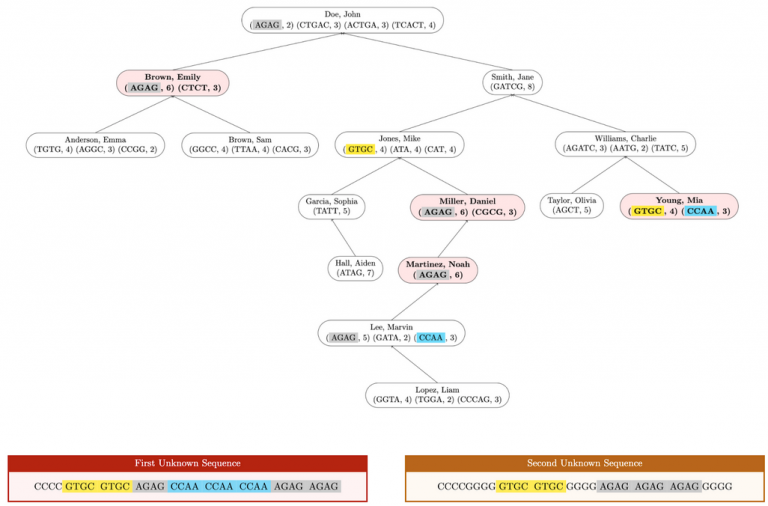
Here is the expected output when testing this method on input1.in. Test assembleTree in the driver before testing this method:

5. getMatchingProfileCount
Find the number of profiles in the BST rooted at rootNode whose marked status matches the isOfInterest parameter. Return this number. If there are no profiles that are marked according to the parameter isOfInterest, return 0.
Recall that Profiles are values stored inside nodes. The Profile.java class contains a boolean instance variable isMarked and getters and setters. By default, all profiles are marked false (false means not of interest).
isMarked is a boolean value: true represents “marked” and false represents unmarked.
In the following diagram, nodes with marked profiles are red and bold. Unmarked profiles are blue and unbolded.
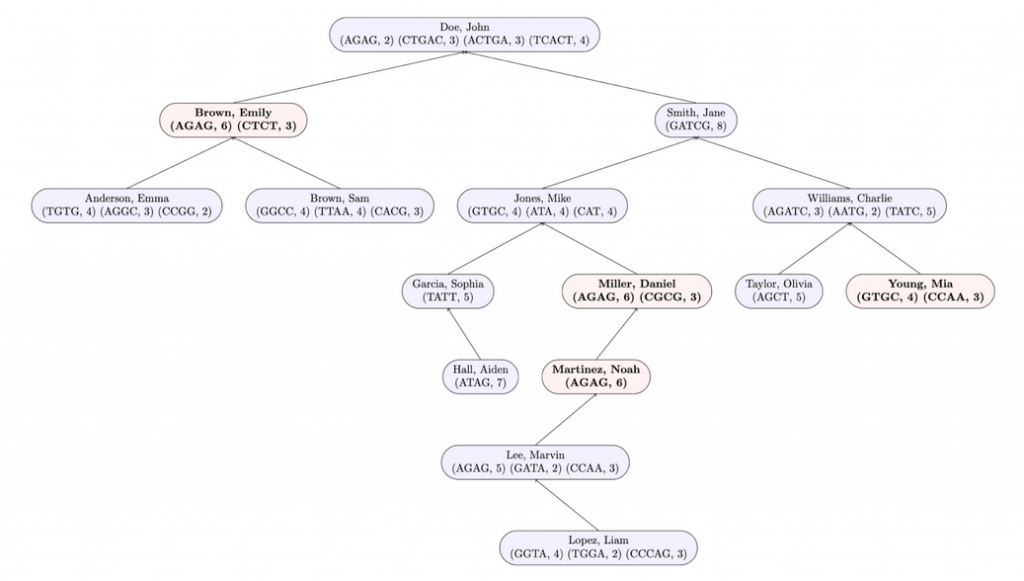
Here is the expected output when testing this method on input1.in. Test assembleTree and markProfilesOfInterest in that order in the driver before testing this method.

6. findUnmarkedPeople
This method uses a level-order traversal to populate and return an array of unmarked profiles in the BST rooted at rootNode.
Use the getMatchingProfileCount method you implemented earlier to get the array length. Create a new array of Strings using the number of unmarked profiles as its length.
Use the provided Queue class to initialize a queue of BSTNodes.
Until the queue is empty:
For each node:
Investigate whether its profile is marked or not. If its profile is not marked, add the PERSON’S NAME (key) to the next available space in your resulting array.
Enqueue the node’s children.
To initialize a queue of BSTNodes, do: Queue<BSTNode<Profile>> queue = new Queue<>();
queue.enqueue(item) enqueues the parameter item into a queue.
queue.dequeue() dequeues an item as discussed in class.
queue.isEmpty() checks if the queue is empty.
Return the resulting array.
Algorithm Example:
The queue starts as empty. Start by enqueuing the root (at position 1).
Dequeue the node at position 1, visit it, and enqueue its left and right children (positions 2 and 3).
Repeat the same procedure on all other nodes until the queue is empty:
Dequeue the node at position 2, visit it, and enqueue its children.
Dequeue the node at position 3, visit it, and enqueue its children.
Lastly, dequeue and visit positions 4 and 5. They are leaves.
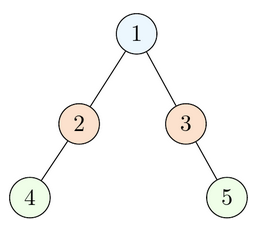
If the nodes at all these positions were marked, they would appear in the resulting array in ascending order of positions. (Note this example above is not a binary search tree but instead just a binary tree, however the algorithm is still the same regardless).
Input File Example: Unmarked profiles are in blue and bold and their corresponding index in the resulting array is next to their name. Marked profiles are in white.
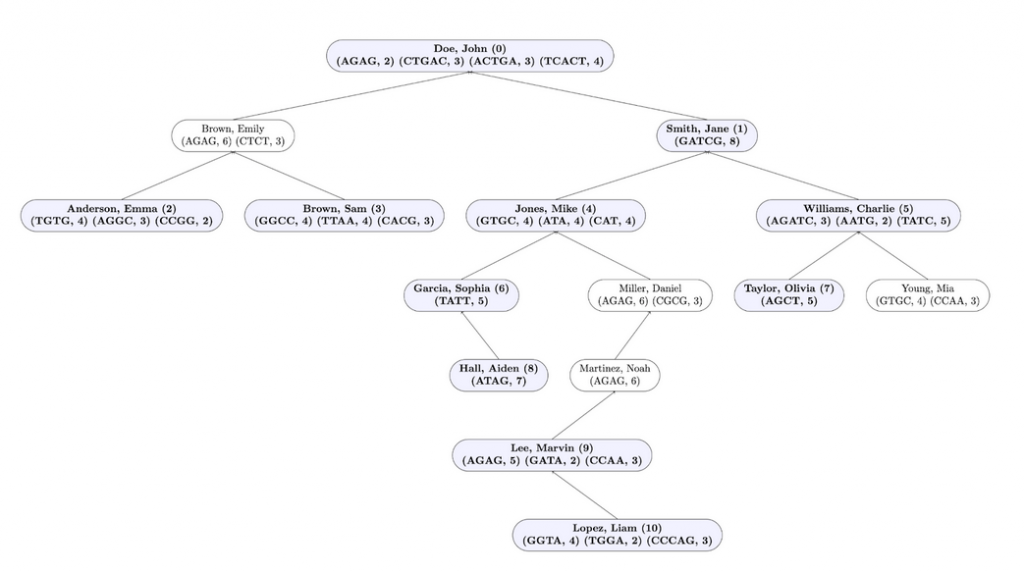
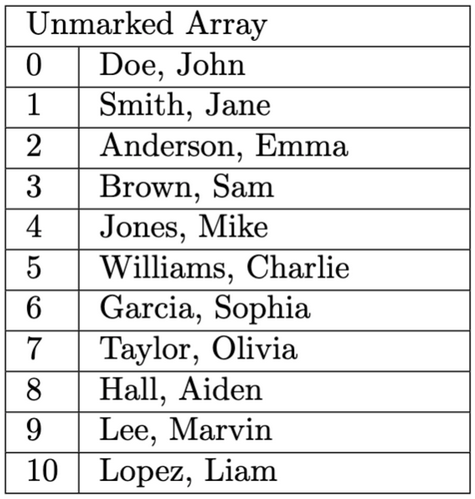
Here is the output for this method when testing this method on input1.in. Test assembleTree and markProfilesOfInterest in that order in the driver before testing this method, and IMPLEMENT getMatchingProfileCount before starting this method.

7. removePerson
This method removes a node whose name matches parameters. This is similar to the BST delete you have seen in class.
Compare full names (“Last, First” including the comma and space) by using the compareTo method on keys as shown in class.
If there are no left or right children, delete the node by setting the parent to null.
If there is only one child, replace the parent link.
If there are two children, use the inorder successor as a replacement for the deleted node.
Note: if the root of the BST changes, rootNode should be updated accordingly.
Here is the output for this method when testing this method on input1.in using “Doe, John”. Test assembleTree in the driver before testing this method, and make sure all methods above are implemented before moving onto the next method. If you have tested other methods, start over.

8. cleanupTree
This method removes nodes containing unmarked profiles from the BST.
Use the findUnmarkedPeople method to get an array of unmarked people.
Use the removePerson method on each unmarked person, passing as parameters the full name of the person.
Here is the output for this method when testing this method on input1.in. Test assembleTree and markProfilesOfInterest in the driver in this order before testing this method, and make sure ALL methods are implemented. Start over if you have called removeProfile in the driver immediately before calling cleanupTree.

Implementation Notes
YOU MAY only update the methods with the WRITE YOUR CODE HERE line.
COMMENT all print statements you have written from DNAProfiles.java
DO NOT add any instance variables to the DNAProfiles class.
DO NOT add any public methods to the DNAProfiles class.
DO NOT add/rename the project or package statements.
DO NOT change the class DNAProfiles name.
YOU MAY add private methods to the DNAProfiles class.
DO NOT use System.exit()
Writing JUnit Tests
You do not have to submit this, this is for you to test your own code.
In the main project folder, there exists a “test” folder next to the “src” folder. This contains a JUnit test class, which can run and test pieces of your code. To compile and run these tests, you must install the Test Runner extension. See the Programming FAQ for more info on VScode extensions.
There are many tests which you can fill in by creating objects, calling methods, and verifying outputs using assert statements. Once you do, remove the fail() statements at the bottom and run.
You are provided with a premade JUnit test class for DNAProfiles. You can finish writing the test methods, by using JUnit test methods (such as assertTrue(), assertFalse(), assertEquals()) in order to test your code.
Tests not running?
First, make sure you’re in the right folder and the tests are implemented. Next, if you have the “Code Runner for Java” or Oracle “Java” extension, make sure you uninstall those extensions.
VSCode Extensions
You can install VSCode extension packs for Java. Take a look at this tutorial. We suggest:
Importing VSCode Project
Download DNAProfiles.zip from Autolab Attachments.
Unzip the file by double clicking.
Open VSCode
Import the folder to a workspace through File > Open Folder
Executing and Debugging
You can run your program through VSCode or you can use the Terminal to compile and execute. We suggest running through VSCode because it will give you the option to debug.
You can hit the triangle “Run Java” button at the top of the Driver.java class, or right click on Driver.java and select “Run Java”
NOTE: if you have DNAProfiles (2) -> DNAProfiles or CS112 -> DNAProfiles in VS Code, open the INNERMOST DNAProfiles through File -> Open Folder.

Before submission
Collaboration policy. Read our collaboration policy on the course syllabus.
Submitting the assignment. Submit DNAProfiles.java separately via the web submission system called Autolab. To do this, go to Autolab and find the correct assignment and submit your Java file.
Getting help
If anything is unclear, don’t hesitate to drop by office hours or post a question on Piazza.
Find instructors and Lead TA office hours here
Find tutors office hours on Canvas -> Tutoring
In addition there is the CSL (Coding and Social Lounge), a community space staffed with student community managers, which are undergraduate students further along the CS major to answer questions.
Inspiration from a Nifty assignment by Brian Yu and David J. Malan.
By Kal Pandit
